1 Basics of R
1.1 Introduction
A <- 2
A # Print A
#> [1] 2
A = 2
A
#> [1] 2
B <- "Halo Semua"
B
#> [1] "Halo Semua"
a<-10 # Space is not sensitive but lettercase is sensitive.
A
#> [1] 2
a
#> [1] 10
# Arithmetic operation
x <- 5
y <- 3
x + y
#> [1] 8
x - y
#> [1] 2
x * y
#> [1] 15
x / y
#> [1] 1.666667
# Logic operation
a <- TRUE
b <- FALSE
a & b
#> [1] FALSE
a | b
#> [1] TRUE
!a
#> [1] FALSE
x <- 5
y <- 3
x > y
#> [1] TRUE
x < y
#> [1] FALSE
x == y
#> [1] FALSE
x >= y
#> [1] TRUE
x <= y
#> [1] FALSE1.2 Types of Objects in R
1.2.1 Vector
a1 <- c(2,4,7,3) # Numeric vector
a2 <- c("one","two","three") # Character vector
a3 <- c(TRUE,TRUE,TRUE,FALSE,TRUE,FALSE) # Logical vector
a1
#> [1] 2 4 7 3
a3[4]
#> [1] FALSE
a2[c(1,3)]
#> [1] "one" "three"
a1[-1]
#> [1] 4 7 3
a1[2:4]
#> [1] 4 7 3
a <- c(1, 2, 3)
b <- c(4, 5, 6)
c <- c(a, b)
c
#> [1] 1 2 3 4 5 6
c[1:3]
#> [1] 1 2 3
d <- a + b
d
#> [1] 5 7 9
a4 <- 1:12
b1 <- matrix(a4,3,4)
b2 <- matrix(a4,3,4,byrow=TRUE)
b3 <- matrix(1:14,4,4)
#> Warning in matrix(1:14, 4, 4): data length [14] is not a
#> sub-multiple or multiple of the number of rows [4]
b1
#> [,1] [,2] [,3] [,4]
#> [1,] 1 4 7 10
#> [2,] 2 5 8 11
#> [3,] 3 6 9 12
b2
#> [,1] [,2] [,3] [,4]
#> [1,] 1 2 3 4
#> [2,] 5 6 7 8
#> [3,] 9 10 11 12
b3
#> [,1] [,2] [,3] [,4]
#> [1,] 1 5 9 13
#> [2,] 2 6 10 14
#> [3,] 3 7 11 1
#> [4,] 4 8 12 2
b2[2,3]
#> [1] 7
b2[1:2,]
#> [,1] [,2] [,3] [,4]
#> [1,] 1 2 3 4
#> [2,] 5 6 7 8
b2[c(1,3),-2]
#> [,1] [,2] [,3]
#> [1,] 1 3 4
#> [2,] 9 11 12
dim(b2)
#> [1] 3 4
m1 <- matrix(c(1, 2, 3, 4, 5, 6), nrow = 2, ncol = 3)
m2 <- matrix(c(7, 8, 9, 10, 11, 12), nrow = 2, ncol = 3)
m3 <- m1 + m2
m3
#> [,1] [,2] [,3]
#> [1,] 8 12 16
#> [2,] 10 14 181.2.2 Factor
levels(d1) <- c("Darah A","Darah AB","Darah B","Darah O")
d1
#> [1] Darah A Darah B Darah AB Darah O
#> Levels: Darah A Darah AB Darah B Darah O
a6 <- c("SMA","SD","SMP","SMA","SMA","SMA","SMA","SMA","SMA","SMA","SMA","SMA","SMA")
d5 <- factor(a6, levels=c("SD","SMP","SMA")) # Skala pengukuran ordinal
levels(d5)
#> [1] "SD" "SMP" "SMA"
d5
#> [1] SMA SD SMP SMA SMA SMA SMA SMA SMA SMA SMA SMA SMA
#> Levels: SD SMP SMA1.2.3 List
a1; b2; d1
#> [1] 2 4 7 3
#> [,1] [,2] [,3] [,4]
#> [1,] 1 2 3 4
#> [2,] 5 6 7 8
#> [3,] 9 10 11 12
#> [1] Darah A Darah B Darah AB Darah O
#> Levels: Darah A Darah AB Darah B Darah O
e1 <- list(a1,b2,d1)
e2 <- list(vect=a1,mat=b2,fac=d1)
e1
#> [[1]]
#> [1] 2 4 7 3
#>
#> [[2]]
#> [,1] [,2] [,3] [,4]
#> [1,] 1 2 3 4
#> [2,] 5 6 7 8
#> [3,] 9 10 11 12
#>
#> [[3]]
#> [1] Darah A Darah B Darah AB Darah O
#> Levels: Darah A Darah AB Darah B Darah O
e2
#> $vect
#> [1] 2 4 7 3
#>
#> $mat
#> [,1] [,2] [,3] [,4]
#> [1,] 1 2 3 4
#> [2,] 5 6 7 8
#> [3,] 9 10 11 12
#>
#> $fac
#> [1] Darah A Darah B Darah AB Darah O
#> Levels: Darah A Darah AB Darah B Darah O
e1[[1]][2]
#> [1] 4
e2$fac
#> [1] Darah A Darah B Darah AB Darah O
#> Levels: Darah A Darah AB Darah B Darah O
e2[2]
#> $mat
#> [,1] [,2] [,3] [,4]
#> [1,] 1 2 3 4
#> [2,] 5 6 7 8
#> [3,] 9 10 11 12
names(e2)
#> [1] "vect" "mat" "fac"1.2.4 Data Frame
Angka <- 11:15
Huruf <- factor(LETTERS[6:10])
f1 <- data.frame(Angka,Huruf)
f1
#> Angka Huruf
#> 1 11 F
#> 2 12 G
#> 3 13 H
#> 4 14 I
#> 5 15 J
f1[1,2]
#> [1] F
#> Levels: F G H I J
f1$Angka
#> [1] 11 12 13 14 15
f1[,"Huruf"]
#> [1] F G H I J
#> Levels: F G H I J
colnames(f1)
#> [1] "Angka" "Huruf"
str(f1)
#> 'data.frame': 5 obs. of 2 variables:
#> $ Angka: int 11 12 13 14 15
#> $ Huruf: Factor w/ 5 levels "F","G","H","I",..: 1 2 3 4 51.3 Data Frame Management
data(iris)
head(iris)
#> Sepal.Length Sepal.Width Petal.Length Petal.Width Species
#> 1 5.1 3.5 1.4 0.2 setosa
#> 2 4.9 3.0 1.4 0.2 setosa
#> 3 4.7 3.2 1.3 0.2 setosa
#> 4 4.6 3.1 1.5 0.2 setosa
#> 5 5.0 3.6 1.4 0.2 setosa
#> 6 5.4 3.9 1.7 0.4 setosa
tail(iris)
#> Sepal.Length Sepal.Width Petal.Length Petal.Width
#> 145 6.7 3.3 5.7 2.5
#> 146 6.7 3.0 5.2 2.3
#> 147 6.3 2.5 5.0 1.9
#> 148 6.5 3.0 5.2 2.0
#> 149 6.2 3.4 5.4 2.3
#> 150 5.9 3.0 5.1 1.8
#> Species
#> 145 virginica
#> 146 virginica
#> 147 virginica
#> 148 virginica
#> 149 virginica
#> 150 virginica
str(iris)
#> 'data.frame': 150 obs. of 5 variables:
#> $ Sepal.Length: num 5.1 4.9 4.7 4.6 5 5.4 4.6 5 4.4 4.9 ...
#> $ Sepal.Width : num 3.5 3 3.2 3.1 3.6 3.9 3.4 3.4 2.9 3.1 ...
#> $ Petal.Length: num 1.4 1.4 1.3 1.5 1.4 1.7 1.4 1.5 1.4 1.5 ...
#> $ Petal.Width : num 0.2 0.2 0.2 0.2 0.2 0.4 0.3 0.2 0.2 0.1 ...
#> $ Species : Factor w/ 3 levels "setosa","versicolor",..: 1 1 1 1 1 1 1 1 1 1 ...1.3.1 R Package
# install.packages("readxl") - code to install R package
library(readxl)
#> Warning: package 'readxl' was built under R version 4.2.3
#install.packages("dplyr")
library(dplyr)
#> Warning: package 'dplyr' was built under R version 4.2.3
#>
#> Attaching package: 'dplyr'
#> The following objects are masked from 'package:stats':
#>
#> filter, lag
#> The following objects are masked from 'package:base':
#>
#> intersect, setdiff, setequal, union
1.3.2 Data Management With dplyr
head(iris)
#> Sepal.Length Sepal.Width Petal.Length Petal.Width Species
#> 1 5.1 3.5 1.4 0.2 setosa
#> 2 4.9 3.0 1.4 0.2 setosa
#> 3 4.7 3.2 1.3 0.2 setosa
#> 4 4.6 3.1 1.5 0.2 setosa
#> 5 5.0 3.6 1.4 0.2 setosa
#> 6 5.4 3.9 1.7 0.4 setosa
irisbaru <- mutate(iris, sepal2 = Sepal.Length + Sepal.Width)
head(irisbaru)
#> Sepal.Length Sepal.Width Petal.Length Petal.Width Species
#> 1 5.1 3.5 1.4 0.2 setosa
#> 2 4.9 3.0 1.4 0.2 setosa
#> 3 4.7 3.2 1.3 0.2 setosa
#> 4 4.6 3.1 1.5 0.2 setosa
#> 5 5.0 3.6 1.4 0.2 setosa
#> 6 5.4 3.9 1.7 0.4 setosa
#> sepal2
#> 1 8.6
#> 2 7.9
#> 3 7.9
#> 4 7.7
#> 5 8.6
#> 6 9.3
irisetosa <- filter(iris, Species=="setosa")
head(irisetosa)
#> Sepal.Length Sepal.Width Petal.Length Petal.Width Species
#> 1 5.1 3.5 1.4 0.2 setosa
#> 2 4.9 3.0 1.4 0.2 setosa
#> 3 4.7 3.2 1.3 0.2 setosa
#> 4 4.6 3.1 1.5 0.2 setosa
#> 5 5.0 3.6 1.4 0.2 setosa
#> 6 5.4 3.9 1.7 0.4 setosa
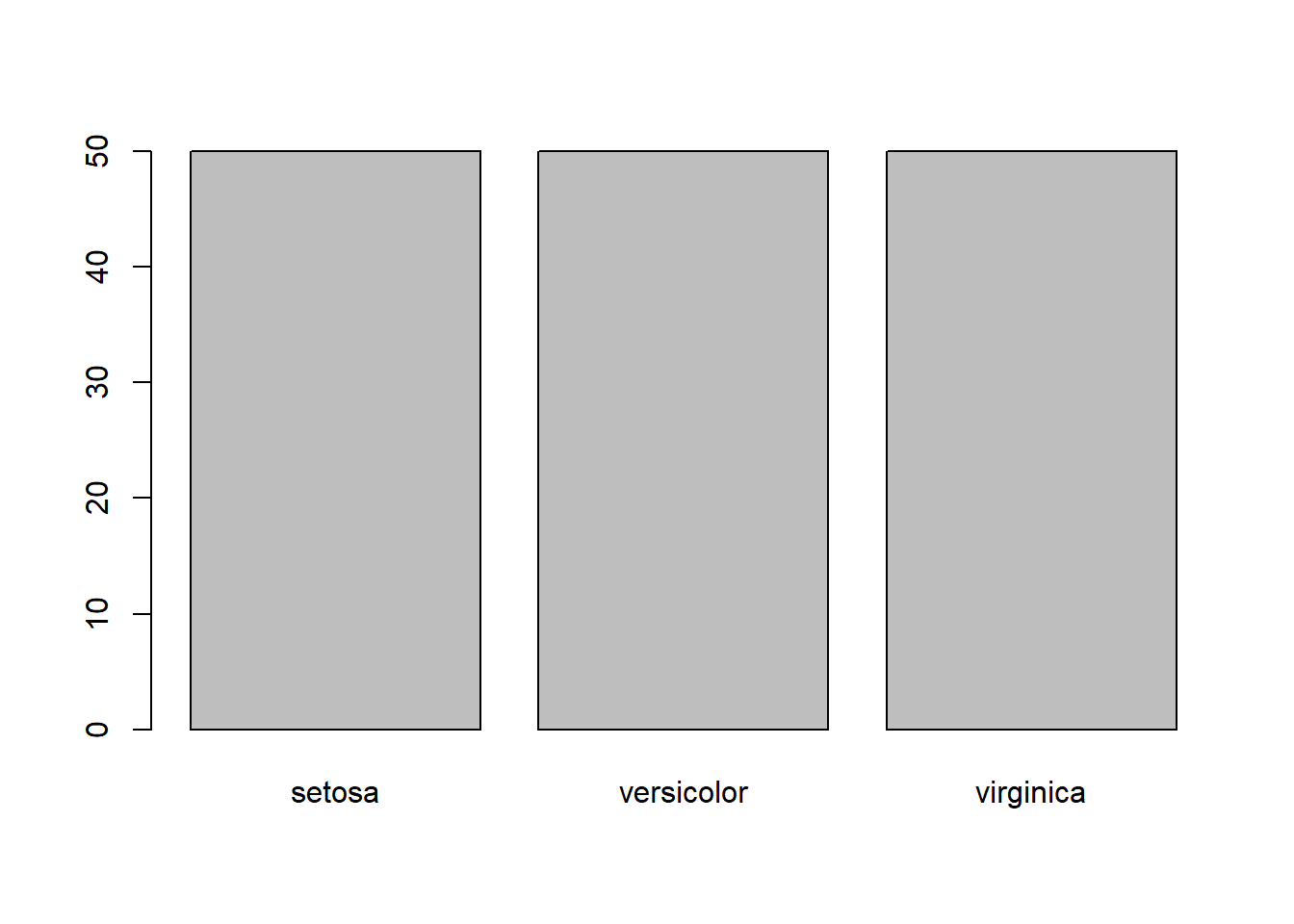
levels(iris$Species)
#> [1] "setosa" "versicolor" "virginica"
irisversicolor <- filter(iris, Species=="setosa"& Petal.Length==1.3)
head(irisversicolor)
#> Sepal.Length Sepal.Width Petal.Length Petal.Width Species
#> 1 4.7 3.2 1.3 0.2 setosa
#> 2 5.4 3.9 1.3 0.4 setosa
#> 3 5.5 3.5 1.3 0.2 setosa
#> 4 4.4 3.0 1.3 0.2 setosa
#> 5 5.0 3.5 1.3 0.3 setosa
#> 6 4.5 2.3 1.3 0.3 setosa
iris3 <- select(iris, Sepal.Length, Species)
head(iris3)
#> Sepal.Length Species
#> 1 5.1 setosa
#> 2 4.9 setosa
#> 3 4.7 setosa
#> 4 4.6 setosa
#> 5 5.0 setosa
#> 6 5.4 setosa
iris4 <- arrange(iris, Petal.Width)
head(iris4)
#> Sepal.Length Sepal.Width Petal.Length Petal.Width Species
#> 1 4.9 3.1 1.5 0.1 setosa
#> 2 4.8 3.0 1.4 0.1 setosa
#> 3 4.3 3.0 1.1 0.1 setosa
#> 4 5.2 4.1 1.5 0.1 setosa
#> 5 4.9 3.6 1.4 0.1 setosa
#> 6 5.1 3.5 1.4 0.2 setosa
iris4 <- arrange(iris, Species, desc(Petal.Width))
head(iris4)
#> Sepal.Length Sepal.Width Petal.Length Petal.Width Species
#> 1 5.0 3.5 1.6 0.6 setosa
#> 2 5.1 3.3 1.7 0.5 setosa
#> 3 5.4 3.9 1.7 0.4 setosa
#> 4 5.7 4.4 1.5 0.4 setosa
#> 5 5.4 3.9 1.3 0.4 setosa
#> 6 5.1 3.7 1.5 0.4 setosa
names(iris4)[1] <- "length"
head(iris4)
#> length Sepal.Width Petal.Length Petal.Width Species
#> 1 5.0 3.5 1.6 0.6 setosa
#> 2 5.1 3.3 1.7 0.5 setosa
#> 3 5.4 3.9 1.7 0.4 setosa
#> 4 5.7 4.4 1.5 0.4 setosa
#> 5 5.4 3.9 1.3 0.4 setosa
#> 6 5.1 3.7 1.5 0.4 setosa